Unidimensional or Multidimensional? Comparing IRT Structural Models
Evidence for a single-ability model with testlet effects
1 Why this matters
Before we can score students on this screener, we need to answer a foundational question: are we measuring one ability or several?
If students’ performance on number-line, magnitude comparison, and other tasks is driven by a single underlying numeracy ability, we should report one overall score. If there are genuinely distinct sub-skills — spatial number sense vs. symbolic comparison, say — then separate profile scores could give teachers more actionable information.
This analysis compares three structural models head-to-head to determine which best describes the data.
2 Three candidate models
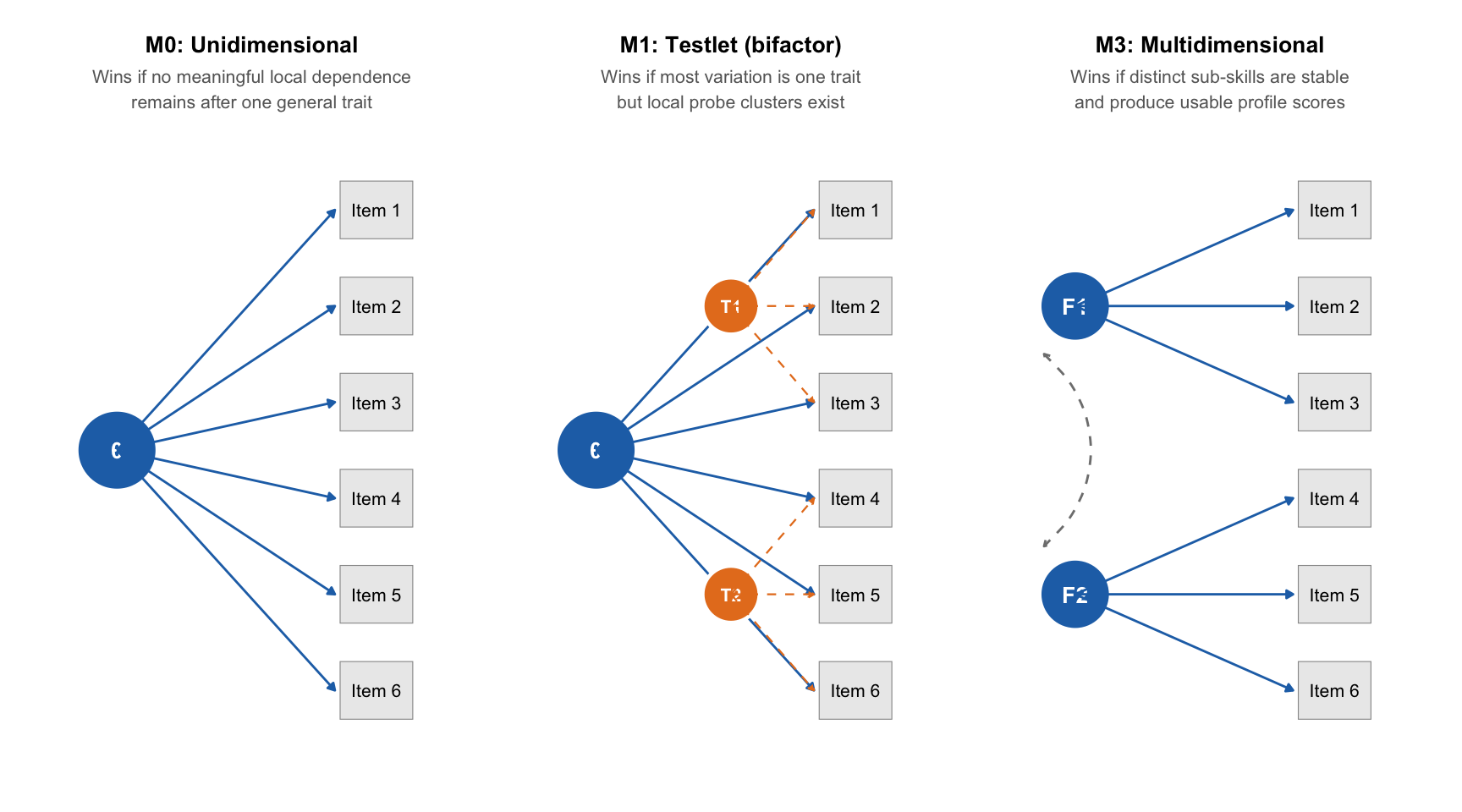
The three models represent increasingly complex accounts of what drives student responses.
| Model | Structure | Wins if... |
|---|---|---|
| M0: Unidimensional | Single latent trait drives all items | No meaningful local dependence or sub-skill structure exists |
| M1: Testlet (bifactor) | One general trait plus testlet factors for probe clusters | Most variation is one trait, but items within a probe are more similar than expected |
| M3: Multidimensional | Multiple correlated latent factors (one per domain) | Distinct sub-skills are stable enough to improve fit and produce usable scores |
2.1 Model equations
All three models share the same item response functions — Rasch for binary items and the generalised partial credit model (GPCM) for number-line items — and differ only in how they parameterise the latent trait(s).
Shared item response functions. For binary items:
\[P(Y_{ij} = 1 \mid \theta_i, b_j) = \text{logit}^{-1}(\theta_i - b_j)\]
For number-line items (3 ordered categories, \(k \in \{0,1,2\}\)):
\[P(Y_{ij} = k \mid \theta_i, b_j, d_{j,\cdot}) \propto \exp\!\left(\sum_{c=1}^{k} (\theta_i - b_j - d_{j,c})\right)\]
where \(Y_{ij}\) is the response of student \(i\) on item \(j\), \(b_j\) is item difficulty, and \(d_{j,c}\) are step thresholds.
M0 — Unidimensional. A single latent trait \(\theta_i\) enters the response functions above directly. No additional structure.
M1 — Testlet (bifactor). A general trait \(\theta_i\) loads on every item. Each probe cluster \(t\) adds a testlet-specific factor \(s_{t,i}\), orthogonal to \(\theta\):
\[\text{logit}\,P(Y_{ij} = 1) = a_j\,\theta_i + a_{S,j}\,s_{t(j),i} - b_j\]
where \(a_j\) and \(a_{S,j}\) are the general and specific loadings. Testlet factors absorb local dependence among items within the same probe without splitting the general ability dimension.
M3 — Multidimensional (correlated factors). Each item loads on a single domain factor \(\theta_{f,i}\):
\[\text{logit}\,P(Y_{ij} = 1) = a_j\,\theta_{f(j),i} - b_j\]
Factor correlations \(\text{Cor}(\theta_f, \theta_g)\) are freely estimated. There is no general factor — all covariation between domains is captured through pairwise correlations.
3 How subtests map to dimensions
M3 with a separate factor for every task failed to converge on the main pooled units, so we collapsed number-line tasks (BNL, UNLC, UNLNC) into a single NL factor and magnitude comparison & missing number tasks (MC, MNA, MNC) into a single SYM_MAG factor. Match quantity and decomposition each keep their own factor. Because each exam group is administered one MQ variant and one DMT variant, a cohort-level unit has 4 factors (NL, SYM_MAG, MQ, DMT); pooled units that combine both exam groups have 6. The diagram below shows all tasks grouped by test family.
Warning: The `label.size` argument of `geom_label()` is deprecated as of ggplot2 3.5.0.
ℹ Please use the `linewidth` argument instead.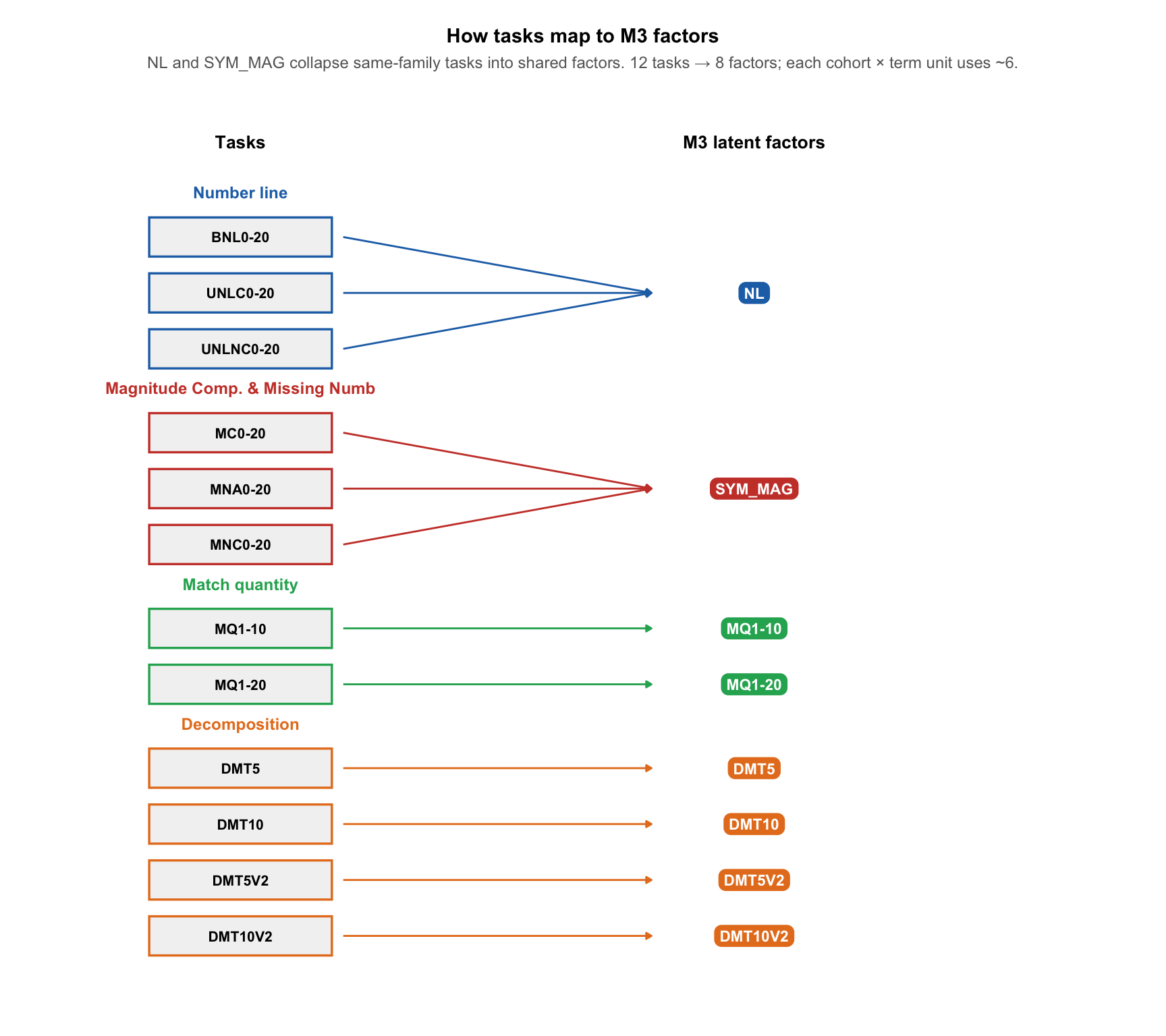
4 What the evidence shows
We evaluated the three models across 12 analysis units (2 terms × 2 sample definitions × 3 cohort scopes), using BIC as the primary selection criterion, residual dependence (Q3) as a secondary check, and school-level bootstrap resampling to test robustness.
4.1 BIC comparison
The analysis was run across 12 units representing every combination of term (Term 1 and Term 4), sample definition (“All students” = every student with at least 5 attempted items in that term; “Stable schools” = only students from the 39 schools present in both terms), and cohort scope (Foundation A, Foundation B, or all foundation students pooled).
The Bayesian Information Criterion penalises model complexity: a lower BIC means the model explains the data better relative to its number of free parameters.
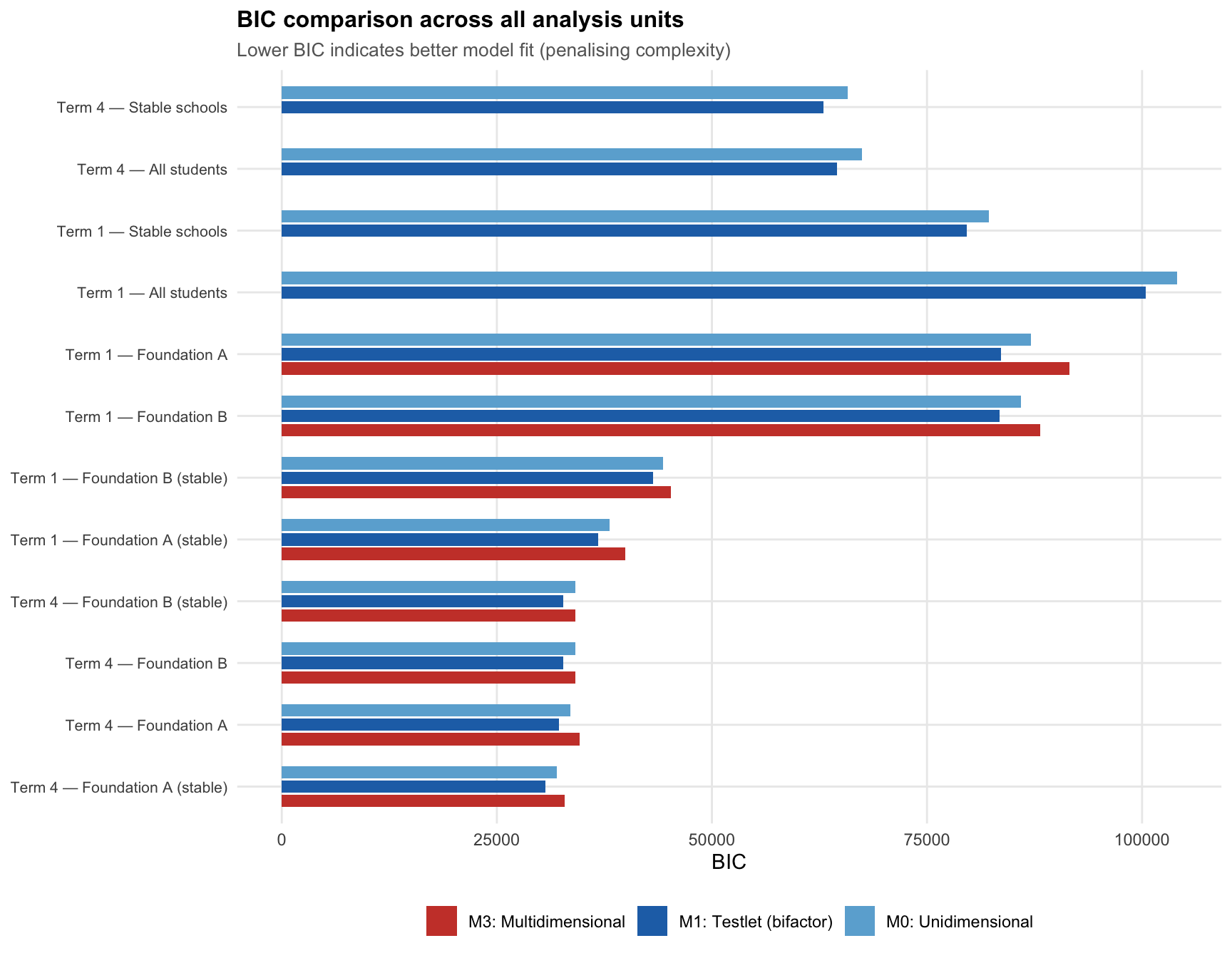
M1 wins in every unit. Across all 12 analysis units — both terms, both sample definitions, all cohorts — the testlet model achieves the lowest BIC. The unidimensional model (M0) consistently ranks second, and the multidimensional model (M3) ranks last.
4.2 Head-to-head BIC differences
To quantify the margin of victory, we examine pairwise BIC differences between models. A negative value in “M1 vs M0” means M1 fits better; a positive value in “M3 vs M1” means M1 fits better than M3.
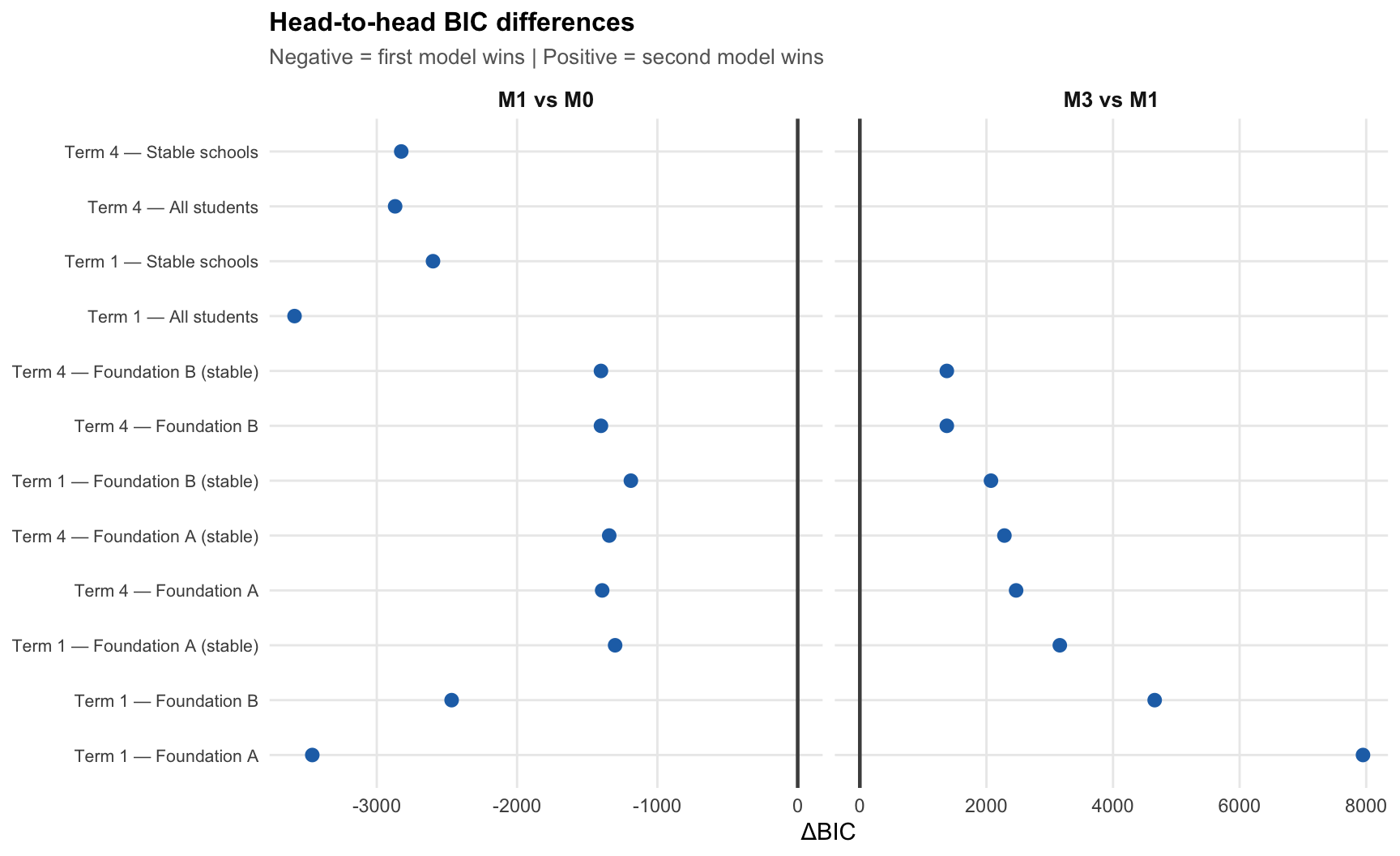
The testlet model beats the unidimensional model by a consistent margin (accounting for local probe dependence), while the multidimensional model performs worse than the testlet model in every unit — often by thousands of BIC points.
4.3 Bootstrap stability
Could these results be an artefact of the particular school sample? To check, we resampled schools 30 times per slice (with replacement) and refit both M1 and M3 on each bootstrap sample. The slices range from approximately 1,100 to 2,500 students; stable-school slices draw from the 39 schools present in both terms.
| Slice | N students | N observations | N schools |
|---|---|---|---|
| Term 1 — All students | 2,514 | 152,068 | — |
| Term 1 — Stable schools | 1,186 | 71,289 | 39 |
| Term 4 — All students | 1,098 | 68,371 | — |
| Term 4 — Stable schools | 1,076 | 67,039 | 39 |
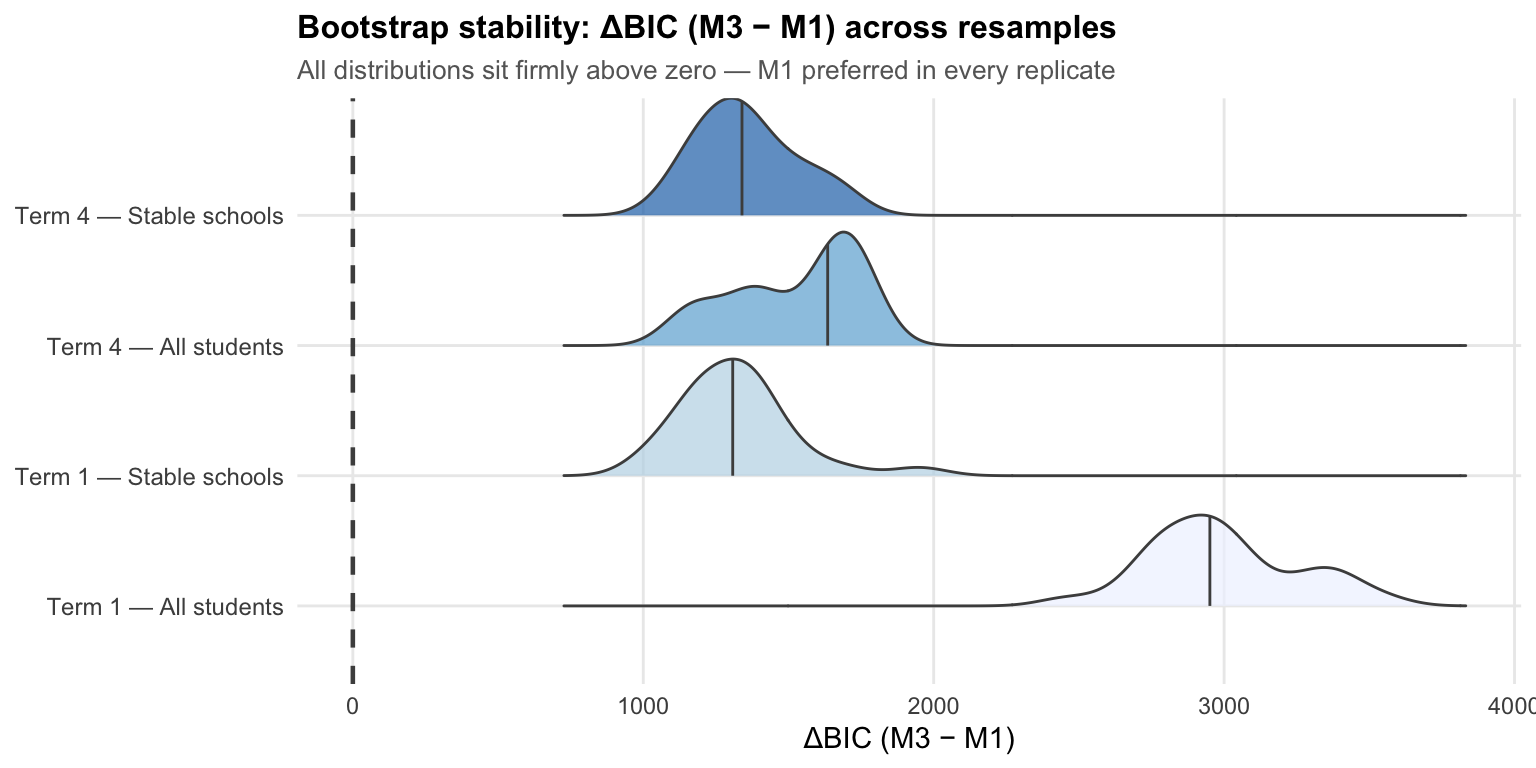
Each ridge shows the distribution of ΔBIC across 30 bootstrap replicates. All four distributions sit entirely above zero — even the most conservative replicate (smallest ΔBIC) favours M1 by over 1,000 BIC points. The wider spread for Term 4 — All students reflects the smaller sample (1,098 students vs 2,514 in Term 1).
4.4 Residual dependence (Q3)
The Q3 statistic measures unexplained local dependence between item pairs after accounting for the latent structure. Lower values indicate the model has adequately captured the correlation structure.
| Scope | Model | Units | Median mean abs(Q3) | Median prop abs(Q3) > threshold |
|---|---|---|---|---|
| By cohort | M0: Unidimensional | 8 | 0.1051 | 0.1424 |
| By cohort | M1: Testlet (bifactor) | 8 | 0.0940 | 0.1146 |
| By cohort | M3: Multidimensional | 8 | 0.1200 | 0.1804 |
| All students | M0: Unidimensional | 4 | 0.1130 | 0.1573 |
| All students | M1: Testlet (bifactor) | 4 | 0.1028 | 0.1355 |
M1 achieves the lowest residual dependence across both scopes: it absorbs local probe effects that M0 leaves unexplained, and does so more efficiently than the multidimensional M3.
5 Conclusion
5.1 M1 (testlet/bifactor) is the preferred structural model
The testlet model wins consistently: it achieves the lowest BIC in all 12 analysis units, demonstrates the lowest residual dependence, and its advantage over both competitors is stable under school-level bootstrap resampling.
Practical implication: the screener should be scored using a single general ability estimate (θ) with testlet effects absorbed into the model. Multidimensional profile scores — separate sub-skill scores for number-line vs. symbolic comparison, for instance — are not viable at current item counts because the multidimensional model fits worse and factor-level precision is insufficient.
Looking ahead: as the item pool grows in future assessment cycles, it will be worth revisiting whether distinct sub-skill profiles become feasible. For now, no profile-level scores should be reported to teachers.
6 Technical appendix
| Unit | BIC M0 | BIC M1 | BIC M3 | Δ M0–best | Δ M1–best | Δ M3–best | Winner |
|---|---|---|---|---|---|---|---|
| Term 1 — Foundation A | 87,094.6 | 83,634.9 | 91,584.3 | 3459.7 | 0 | 7949.4 | M1_1d_testlets |
| Term 1 — Foundation B | 85,936.3 | 83,469.7 | 88,127.7 | 2466.6 | 0 | 4658.0 | M1_1d_testlets |
| Term 1 — Foundation A (stable) | 38,089.5 | 36,788.2 | 39,947.5 | 1301.2 | 0 | 3159.2 | M1_1d_testlets |
| Term 1 — Foundation B (stable) | 44,334.2 | 43,145.2 | 45,216.9 | 1189.0 | 0 | 2071.7 | M1_1d_testlets |
| Term 4 — Foundation A | 33,577.8 | 32,184.3 | 34,653.1 | 1393.6 | 0 | 2468.9 | M1_1d_testlets |
| Term 4 — Foundation B | 34,125.4 | 32,723.3 | 34,097.5 | 1402.2 | 0 | 1374.3 | M1_1d_testlets |
| Term 4 — Foundation A (stable) | 31,965.5 | 30,622.2 | 32,906.2 | 1343.3 | 0 | 2284.0 | M1_1d_testlets |
| Term 4 — Foundation B (stable) | 34,125.4 | 32,723.3 | 34,097.5 | 1402.2 | 0 | 1374.3 | M1_1d_testlets |
| Term 1 — All students | 104,043.8 | 100,457.6 | NA | 3586.2 | 0 | NA | M1_1d_testlets |
| Term 1 — Stable schools | 82,192.0 | 79,592.7 | NA | 2599.3 | 0 | NA | M1_1d_testlets |
| Term 4 — All students | 67,401.5 | 64,532.3 | NA | 2869.2 | 0 | NA | M1_1d_testlets |
| Term 4 — Stable schools | 65,799.3 | 62,973.6 | NA | 2825.6 | 0 | NA | M1_1d_testlets |
| Unit | Model | Mean abs(Q3) | Max abs(Q3) | Prop abs(Q3) > threshold |
|---|---|---|---|---|
| Term 1 — Foundation A | M0: Unidimensional | 0.0953 | 0.9828 | 0.1257 |
| Term 1 — Foundation A | M1: Testlet (bifactor) | 0.0856 | 0.9977 | 0.1145 |
| Term 1 — Foundation A | M3: Multidimensional | 0.1220 | 1.0000 | 0.1949 |
| Term 1 — Foundation A (stable) | M0: Unidimensional | 0.0979 | 1.0000 | 0.1228 |
| Term 1 — Foundation A (stable) | M1: Testlet (bifactor) | 0.0890 | 0.7938 | 0.1050 |
| Term 1 — Foundation A (stable) | M3: Multidimensional | 0.1216 | 0.7943 | 0.1829 |
| Term 4 — Foundation A | M0: Unidimensional | 0.1047 | 0.9870 | 0.1437 |
| Term 4 — Foundation A | M1: Testlet (bifactor) | 0.0931 | 0.9792 | 0.1092 |
| Term 4 — Foundation A | M3: Multidimensional | 0.1258 | 0.9880 | 0.1841 |
| Term 4 — Foundation A (stable) | M0: Unidimensional | 0.1074 | 0.9964 | 0.1471 |
| Term 4 — Foundation A (stable) | M1: Testlet (bifactor) | 0.0958 | 0.9943 | 0.1146 |
| Term 4 — Foundation A (stable) | M3: Multidimensional | 0.1265 | 0.9974 | 0.1825 |
| Term 1 — Foundation B | M0: Unidimensional | 0.1017 | 1.0000 | 0.1335 |
| Term 1 — Foundation B | M1: Testlet (bifactor) | 0.0952 | 0.8873 | 0.1274 |
| Term 1 — Foundation B | M3: Multidimensional | 0.1073 | 0.9454 | 0.1503 |
| Term 1 — Foundation B (stable) | M0: Unidimensional | 0.1124 | 0.9958 | 0.1645 |
| Term 1 — Foundation B (stable) | M1: Testlet (bifactor) | 0.1068 | 0.9619 | 0.1529 |
| Term 1 — Foundation B (stable) | M3: Multidimensional | 0.1184 | 0.9769 | 0.1784 |
| Term 4 — Foundation B | M0: Unidimensional | 0.1054 | 1.0000 | 0.1424 |
| Term 4 — Foundation B | M1: Testlet (bifactor) | 0.0940 | 0.9554 | 0.1147 |
| Term 4 — Foundation B | M3: Multidimensional | 0.1030 | 0.9880 | 0.1369 |
| Term 4 — Foundation B (stable) | M0: Unidimensional | 0.1054 | 1.0000 | 0.1424 |
| Term 4 — Foundation B (stable) | M1: Testlet (bifactor) | 0.0940 | 0.9554 | 0.1147 |
| Term 4 — Foundation B (stable) | M3: Multidimensional | 0.1030 | 0.9880 | 0.1369 |
| Term 1 — All students | M0: Unidimensional | 0.1268 | 1.0000 | 0.1940 |
| Term 1 — All students | M1: Testlet (bifactor) | 0.1180 | 1.0000 | 0.1808 |
| Term 1 — Stable schools | M0: Unidimensional | 0.1125 | 1.0000 | 0.1542 |
| Term 1 — Stable schools | M1: Testlet (bifactor) | 0.1049 | 1.0000 | 0.1445 |
| Term 4 — All students | M0: Unidimensional | 0.1125 | 1.0000 | 0.1561 |
| Term 4 — All students | M1: Testlet (bifactor) | 0.0997 | 0.9941 | 0.1237 |
| Term 4 — Stable schools | M0: Unidimensional | 0.1136 | 1.0000 | 0.1584 |
| Term 4 — Stable schools | M1: Testlet (bifactor) | 0.1007 | 0.9938 | 0.1266 |
| Slice | Planned reps | Usable reps | Median ΔBIC | P10 ΔBIC | P90 ΔBIC |
|---|---|---|---|---|---|
| Term 1 — All students | 30 | 30 | 2,951.0 | 2,726.8 | 3,379.8 |
| Term 1 — Stable schools | 30 | 28 | 1,308.0 | 1,130.8 | 1,524.9 |
| Term 4 — All students | 30 | 17 | 1,635.0 | 1,194.8 | 1,743.6 |
| Term 4 — Stable schools | 30 | 14 | 1,339.9 | 1,190.2 | 1,585.2 |
| all_slices | 120 | NA | 1,536.3 | 1,183.4 | 3,062.1 |
Positive values throughout confirm M1 is preferred over M3 in every bootstrap replicate. The P10 column shows even the most conservative replicate favours M1 by a large margin.
Three-model comparison run
- Run ID:
foundation_parallel_20260206 - Source path:
../../analysis/structure_model_comparison_v4/outputs/runs/foundation_parallel_20260206
Full-N robustness run (bootstrap)
- Run ID:
foundation_fulln_boot4_prod_20260209 - Source path:
../../analysis/structure_model_comparison_v4/outputs/runs/foundation_fulln_boot4_prod_20260209
| Run | Status | Converged | Count |
|---|---|---|---|
| Full-N robustness | ok | TRUE | 8 |
| Three-model comparison | ok | TRUE | 32 |
| Three-model comparison | timed_out | FALSE | 4 |
The structural model selection follows a layered decision protocol:
Primary — BIC (Bayesian Information Criterion): the model with the lowest BIC across the majority of analysis units is preferred. BIC penalises free parameters, guarding against overfitting.
Secondary — Residual dependence (Q3): Yen’s Q3 statistic measures unexplained pairwise item dependence after conditioning on the latent structure. Lower mean |Q3| and a lower proportion of item pairs exceeding the threshold (|Q3| > 0.2) indicate better local-dependence absorption.
Robustness — School-level bootstrap: schools are resampled with replacement (30 replicates per slice) and both M1 and M3 are refit. If the BIC preference reverses in a meaningful proportion of replicates, the result is considered unstable.
Profile gate (secondary): even if a multidimensional model won on fit, profile scores are only reported to teachers if factor-level measurement precision exceeds minimum thresholds (determinacy > 0.80 for teacher-facing scores, > 0.70 for internal use). This gate is currently not met for any M3 factor.
“Teacher profile pass” and “Internal profile pass” indicate whether M3’s factor-level precision exceeds minimum thresholds for reporting profile scores (determinacy > 0.80 for teacher-facing, > 0.70 for internal use). All units show FALSE — no M3 factor achieves sufficient precision for profile scores.
| Unit | BIC winner | BIC value | Teacher profile pass | Internal profile pass |
|---|---|---|---|---|
| Term 1 — Foundation A | M1_1d_testlets | 83,634.9 | FALSE | FALSE |
| Term 1 — Foundation A (stable) | M1_1d_testlets | 36,788.2 | FALSE | FALSE |
| Term 4 — Foundation A | M1_1d_testlets | 32,184.3 | FALSE | FALSE |
| Term 4 — Foundation A (stable) | M1_1d_testlets | 30,622.2 | FALSE | FALSE |
| Term 1 — Foundation B | M1_1d_testlets | 83,469.7 | FALSE | FALSE |
| Term 1 — Foundation B (stable) | M1_1d_testlets | 43,145.2 | FALSE | FALSE |
| Term 4 — Foundation B | M1_1d_testlets | 32,723.3 | FALSE | FALSE |
| Term 4 — Foundation B (stable) | M1_1d_testlets | 32,723.3 | FALSE | FALSE |
| Term 1 — All students | M1_1d_testlets | 100,457.6 | FALSE | FALSE |
| Term 1 — Stable schools | M1_1d_testlets | 79,592.7 | FALSE | FALSE |
| Term 4 — All students | M1_1d_testlets | 64,532.3 | FALSE | FALSE |
| Term 4 — Stable schools | M1_1d_testlets | 62,973.6 | FALSE | FALSE |